|
Rate estimates (for Phasmatodea and Zygentoma) and model comparison indicate that the asexual lineages likely mutate several times faster than their sexual relatives. 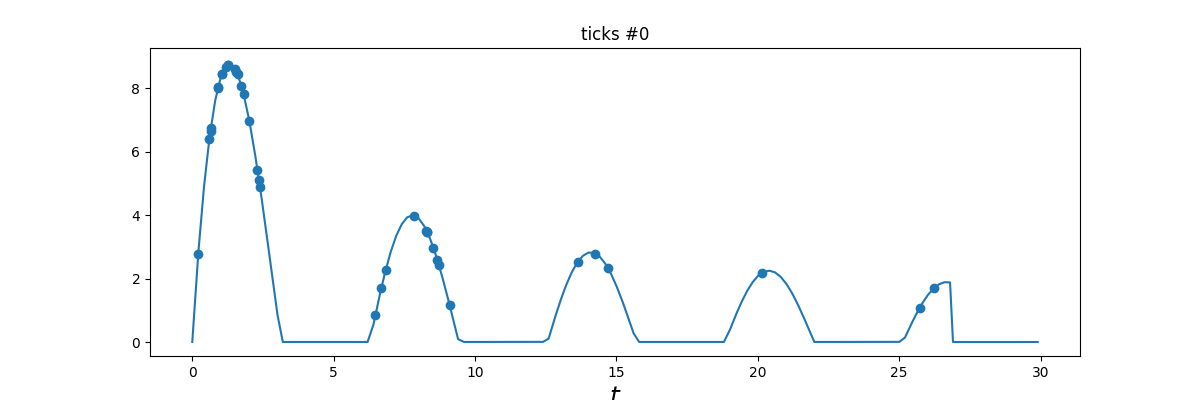
We finally test our procedure on both simulated and recently studied hexapod transcriptome data (Brandt et al.), where each asexual lineage pairs with its closest related sexual lineage. Haggstrom represents a specific, individual, material embodiment of a distinct intellectual or artistic creation found in Indiana State Library. Event queue sampling and probability magnifiers are used to improve the computational efficiency when the number of events is large. Each time you run the Poisson process, it will produce a different sequence of random outcomes as per some probability distribution. The item Sequential tests for exponential populations and poisson processes, Gus W. We calculate probability Pr( B −1 |ɑ) based on the identified events (during “ ɑ→B −1”) from pairwise sequence alignment to implement Pr( B −1| B 1) calculation (Monte Carlo integration). The corresponding retrospective process (reversing the prospective process) facilitates developing algorithms to calculate the marginal probability (Monte Carlo integration) and sample ɑ (given B 1). asymptotic test for incorporating the historical control data for the. DOWNLOADS GCalc Testing Poisson Process - enter up to 14 sets of value. 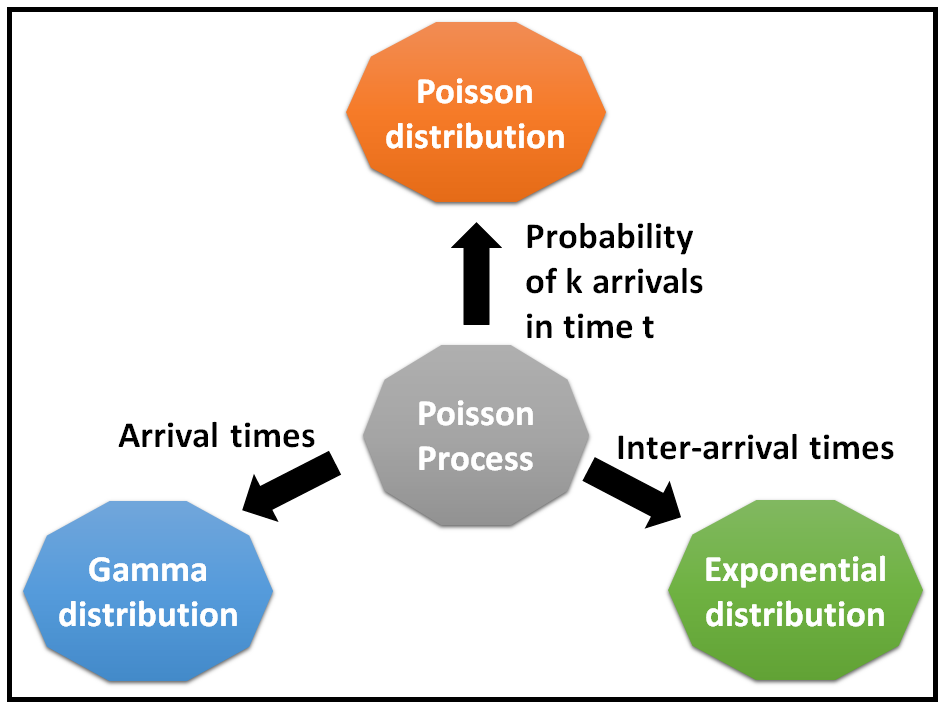
We extend the popular Jukes–Cantor evolution model and calculate the probability of an orthologous nucleotide sequence set, where the sequence evolution follows the (prospective) Poisson process with the overall event rate λ prorated among mutation types (nucleotide/codon substitution, insertion, and deletion) and sites along each sequence. Assuming that Y for a given experiment follows a Poisson distribution with. This on-line calculator plots poisson distribution of the random variable X 19.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
